Elzo de Wit
Genome function and dynamics
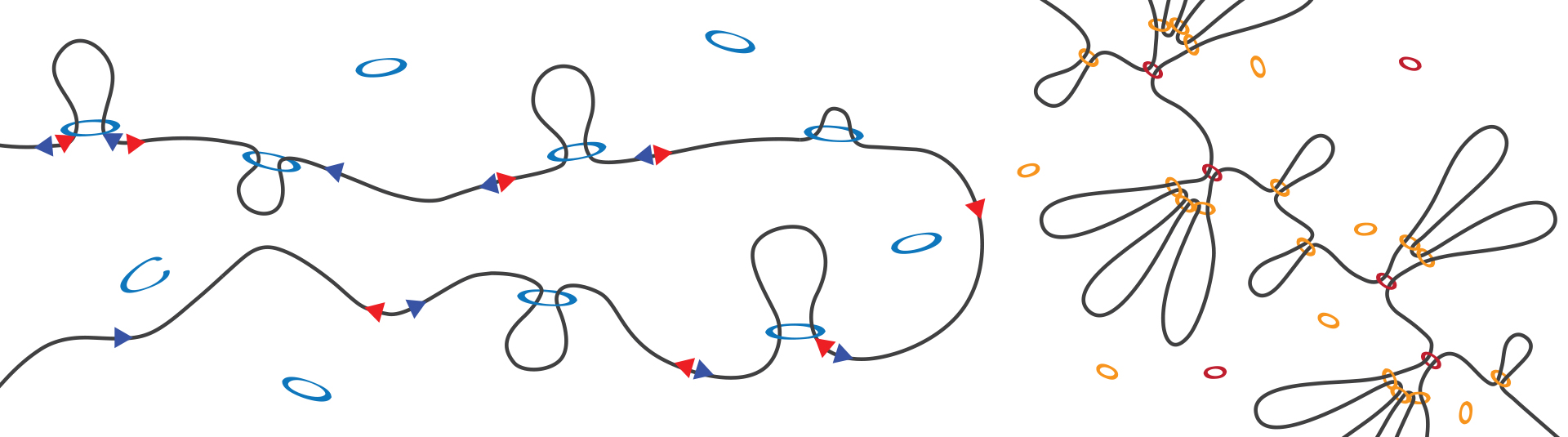
Storage of our DNA inside the nucleus is a formidable task. When stretched out our genome measures 2 meters in length, but it has to fit into a nucleus that is one hundredth of a millimeter in diameter. To achieve this the genome is very efficiently folded.
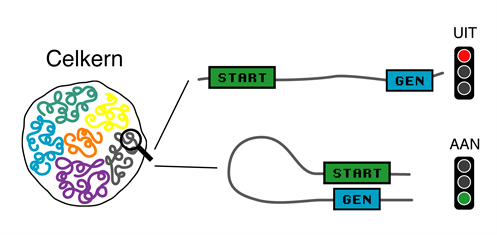
It has become clear that the 3D organization of the genome plays an important role in the regulation of genes. We use state-of-the-art tools such as acute protein depletion, RNA sequencing and Hi-C to study the interplay between genome folding and gene expression and the regulatory factors involved.