T cell dysfunction is a key problem in cancer, enabling not only tumorigenesis but also causing resistance to immunotherapy. We recently obtained an ERC Advanced grant called “ReverT”. This is a genome-wide CRISPR-Cas9 screening program to identify genes, ablation of which reverses dysfunction in primary T cells. This program has just been launched. Our proof-of-concept results uncovered several “nodal” factors, operating in several seemingly different dysfunction settings, which may thus in fact be linked. We will use a collection of adoptive cell transfer mouse and human tumor models for validation and mechanistic characterization, as well as primary human T cells and patient-derived tumor fragments. Lastly, we will translate our findings to a preclinical setting, aiming to achieve more durable clinical responses. Our first series of T cell dysfunction screens has recently been published in Cancer Cell (highlights in Cancer Discovery and ACIR). Furthermore, you can mine your favorite gene in our T cell dysfunction screening datasets.
A functional immune defense against cancer depends on many parameters, including specific physical and dynamic contacts between cytotoxic CD8+ T cells and tumor cells, triggering numerous signaling events between and within these cell types. Whereas disruptions in T cell:tumor cell interactions contribute to immune
evasion and cancer, pharmacologic intervention of interactions like PD-1:PD-L1 has significant clinical benefit. Interactions between CD8+ T cells and other immune cells have been well-characterized, but we lack a detailed map of functional interactions between CD8+ T cells and tumor cells. We’ve set out to perform high-resolution mapping and functional dissection of the T cell:tumor cell interactome, to provide new mechanistic insight identify novel pharmacologic targets to improve immunotherapy.
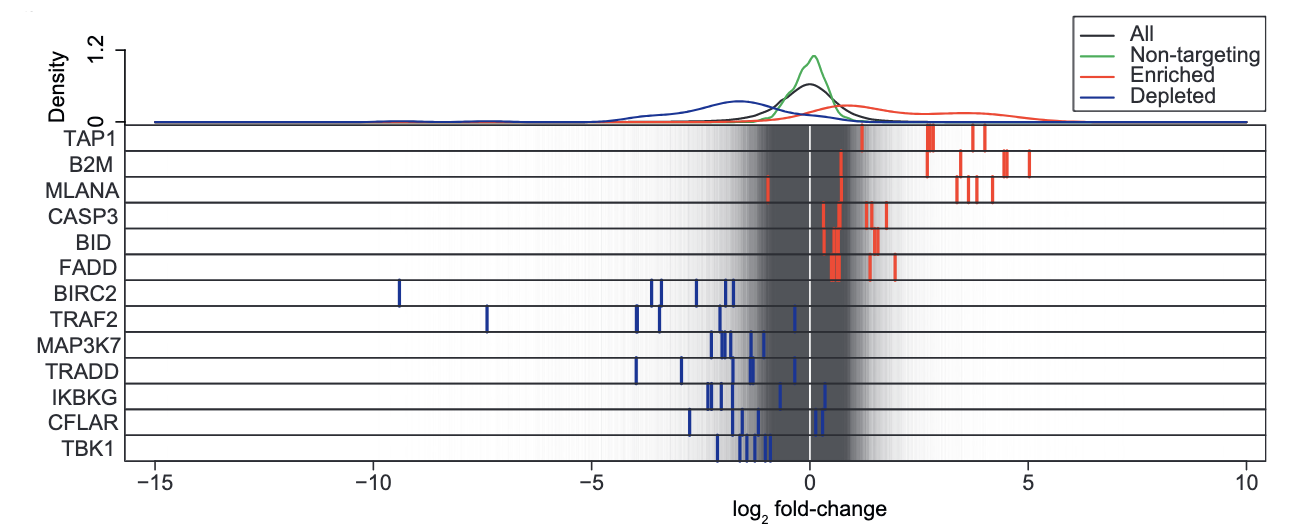
We’ve built several in vitro and in vivo systems to uncover critical mediators of the sensitivity of tumor cells to cytotoxic T cells. For example, we set out to break intrinsic resistance of melanoma to T cell killing. A genome-wide CRISPR/Cas9 screen uncovered several hits mapping to the tumor necrosis factor (TNF) pathway. Clinically, TNF antitumor activity is limited in tumors of ICB non-responders, correlating with its low abundance. Taking advantage of the genetic screen, we demonstrated that ablation of the top hit, TRAF2, lowers the TNF cytotoxicity threshold in tumors by redirecting TNF signaling to favor RIPK1-dependent apoptosis. Our results suggest that selective reduction of the TNF cytotoxicity threshold increases the susceptibility of tumors to immunotherapy (Vredevoogd, Kuilman et al, Cell 2019).
More recently, we focused on common tumor escape mechanisms for immunotherapy, particularly deficiencies in antigen presentation, diminishing adaptive CD8+ T cell antitumor activity. We performed parallel genome-wide CRISPR-Cas9 knockout screens under NK and CD8+ T cell pressure. All components, RNF31, RBCK1, and SHARPIN, of the linear ubiquitination chain assembly complex (LUBAC) were identified as top hits. Genetic and pharmacologic ablation of RNF31, an E3 ubiquitin ligase, strongly sensitized cancer cells to NK and CD8+ T cell killing. This occurred in a tumor necrosis factor (TNF)- dependent manner, causing loss of A20 and non-canonical IKK complexes from TNF receptor complex I. A small-molecule RNF31 inhibitor sensitized tumor organoids to TNF and greatly enhanced bystander killing of MHC antigen-deficient tumor cells. These results merit exploration of RNF31 inhibition as a clinical pharmacological opportunity for immunotherapy-refractory cancers (Zhang, Kong, Ligtenberg et al., Cell Rep. Med., 2022).
We’re also taking more generic approaches to tackle immune resistance. For example, to mimic recurrent T cell attack, we chronically exposed a panel of (patient-derived) melanoma cell lines to cytotoxic T cells. This led to strong enrichment of a pre-existing cell population that exhibited immune resistance in vitro and in mice. These fractions showed high expression of NGFR, were maintained stably, and were found to be present in patients’ melanomas prior to treatment. Remarkably, these cells exhibited multidrug-resistance to other therapies including BRAF + MEK inhibition, suggesting that they exist in a stable and distinct cellular state (Boshuizen et al., Nature Comm. 2020).
Despite our understanding of downstream signaling events of IFNγ, little is known about regulation of its receptor (IFNγ-R1). With an unbiased genome-wide CRISPR/Cas9 screen we identify STUB1 as an E3 ubiquitin ligase for IFNγ-R1 in complex with its signal-relaying kinase JAK1. STUB1 mediates ubiquitination- dependent proteasomal degradation of IFNγ-R1/JAK1 complex through IFNγ-R1K285 and JAK1 K249. Conversely, STUB1 inactivation amplifies IFNγ signaling, sensitizing tumor cells to cytotoxic T cells in vitro. Anti-PD-1 response was increased in heterogenous tumors comprising both wildtype and STUB1-deficient cells, but not full STUB1 knockout tumors. These results uncover STUB1 as a critical regulator of IFNγ-R1 and highlight the context-dependency of STUB1-regulated IFNγ signaling for ICB outcome (Apriamashvili, Vredevoogd et al., Nature Comm. 2022).
The objectives outlined above illustrate that a central goal of our laboratory is to translate our findings to the benefit of the patient, taking advantage of our comprehensive cancer institute. NKI-AVL has an excellent track record in translational and investigator-initiated studies. We have several ongoing collaborations with our clinicians, facilitating translation of our laboratory findings (therapeutic targets, prognostic and predictive biomarkers) to the oncology clinic, for melanoma, lung cancer, sarcoma and other tumor indications.