“Recent years have been a bit like a revolution”, explains NKI group leader Benjamin Rowland. “We have learned that a small group of protein complexes is responsible for a large part of the DNA-shaping processes inside cells. Keep in mind that DNA is the code of life, so the regulation of DNA is not a trivial matter.” These protein complexes include the SMC molecular motors known as cohesin and condensin. By shaping DNAs, they control some of the most basic processes in our cells, ranging from cell division, to the regulation of our genes, and the repair of broken DNAs. They perform similar roles across the tree of life, from bacteria to humans. Benjamin: “Now we know that these complexes are so important, the next question is how they then do what they do.”
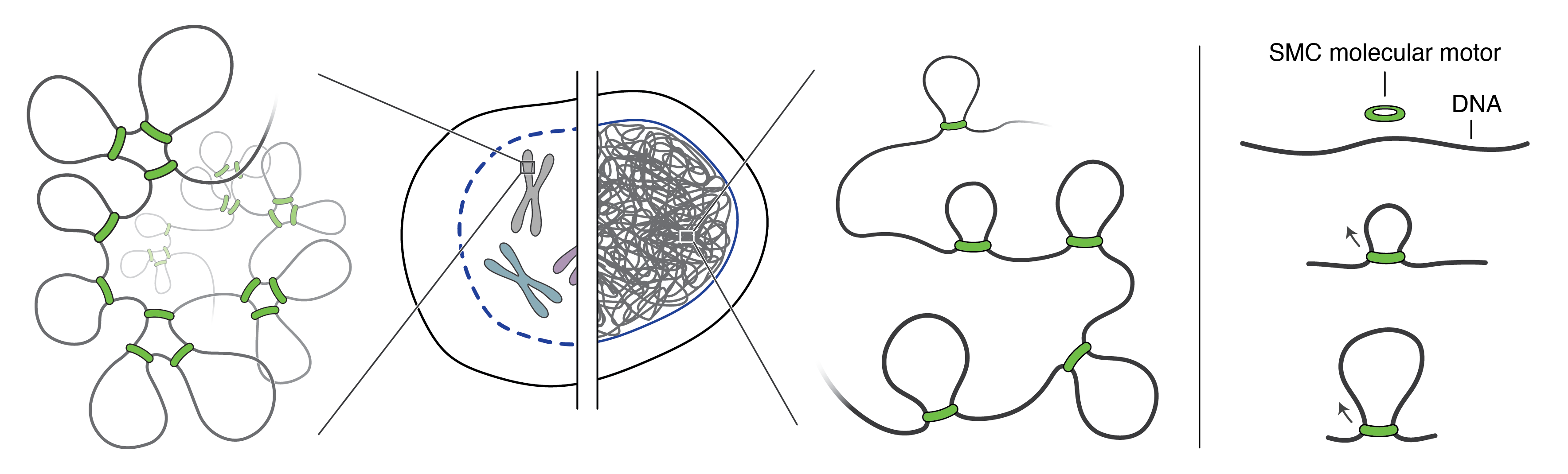
Figure 1. Meters of DNA fit into our tiny cells. This DNA is folded into loops by ring-shaped SMC molecular motors. Click to enlarge.
These molecular motors look like rings, making loops in our DNA (see figure). This might sound simple until you try to describe this process at a molecular level. As they do, scientists try to understand processes by tackling small questions one at a time. To make sense of the bits of knowledge they gather, they try to put together a model of how these motors work. Over the years this has led to various models with appealing names such as ‘swing and clamp’, ‘hold and feed’, and ‘segment capture’. Benjamin: “Each of these models has its own merit, as it explains an important part of the mechanism. I was at a Titisee conference together with three other experts in the field, and we felt that it would be good to make the point that there is a potential unified model that incorporates central aspects of each of these models.”
Earlier that year, Rowland and his PhD student Roel Oldenkamp had noticed something peculiar. Two great papers had just appeared online, but each paper came to an opposite conclusion. One study found that complexes, while building loops, could pass enormous obstacles on the DNA without ever needing to open the ring. The loop evidently was not inside the ring, but had to be held onto by the outer parts of the ring. The other study provided a wealth of insights into the different conformational states of the complex, but one of their conclusions was that the loop surely had to be inside the ring. So what was going on?
“We reasoned that all the data presumably was correct, but that there had to be an alternative interpretation of some of the data that would explain everything. Roel and I spent many hours discussing and drawing together – all online, as it was the height of the Corona pandemic – trying to solve the incongruity. And we did it: we found a solution, and came up with a model that resolved this contradiction.”
Inspired by the incredible progress that had been made by the field in recent years, and by the realization that there no longer was any contradiction, Rowland and his colleagues Cees Dekker, Christian Haering and Jan-Michael Peters sat down together, and racked their brains to make sense of how condensin and cohesin might work. Together, they propose a model called “reel and seal" for how these molecular motors could build DNA loops (see figure and movie). They wrote up their ideas, which they have now published in Science.
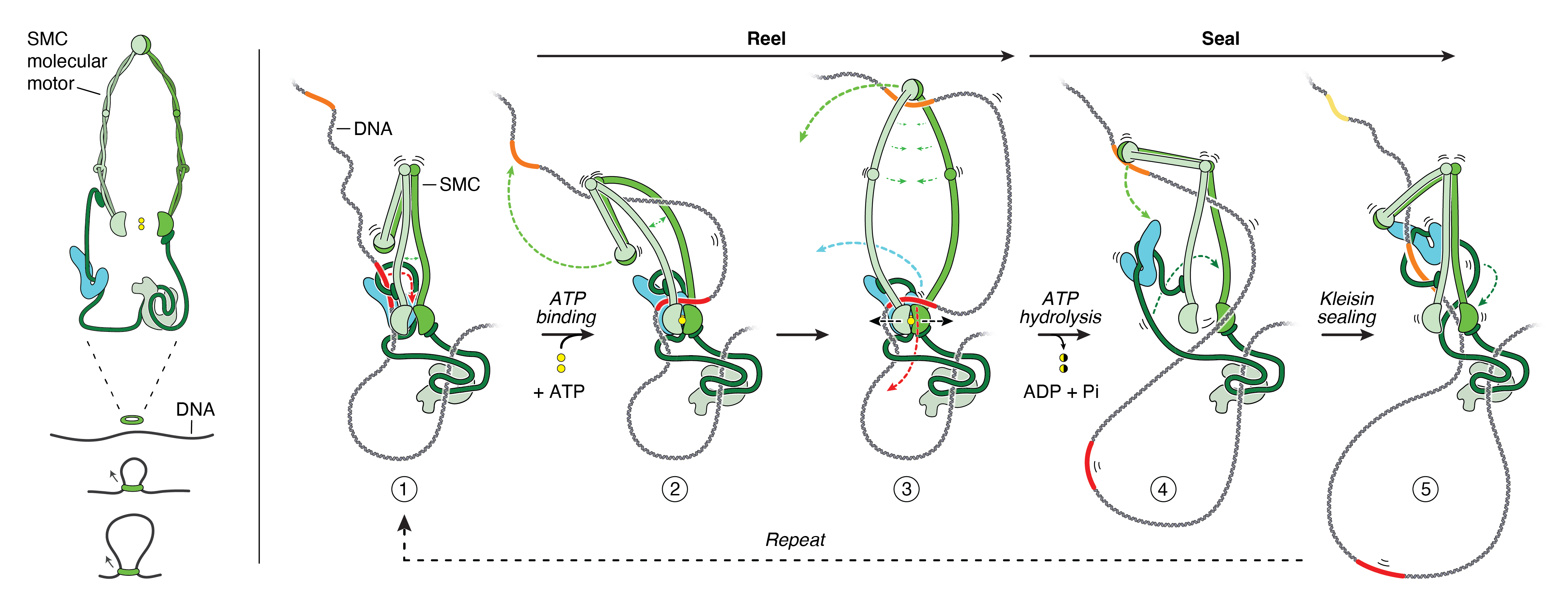
Figure 2. The proposed “reel-and-seal” mechanism for DNA loop extrusion by SMC molecular motors. Click to enlarge.